Introduction
In this article, we will go through the tutorial for Naive Bayes classification in Python Sklearn. We will understand what is Naive Bayes algorithm and proceed to see an end-to-end example of implementing the Gaussian Naive Bayes classifier in Sklearn with a dataset.
What is Naive Bayes Algorithm?
Naive Bayes is a simple yet powerful probabilistic classification model in machine learning that takes inspiration from Bayes Theorem.
Bayes theorem is a formula that gives a conditional probability of an event A taking place provided another event B has already occurred. Its formula can be written as –
\(P(A|B) = \frac{P(B|A).P(A)}{P(B)}\)where
- A and B are two events
- P(A|B) is the probability of event A given event B has already occurred.
- P(B|A) is the probability of event B given event A has already occurred.
- P(A) is the independent probability of A
- P(B) is the independent probability of B
Now, this Bayes theorem can be repurposed to create a classification model as follows –
\(P(y|X) = \frac{P(X|y).P(X)}{P(y)}\)where
- X = x1,x2,x3,.. xN are list of independent predictors
- y is the class label
- P(y|X) is the probability of label y given the predictors X
The above equation can be expanded as
\(P(y|x1,x2,x3..xN) = \frac{P(x1|y).P(x2|y).P(x3|y)… P(xN|y).P(y)}{P(x1).P(x2).P(x3)…P(xN)}\)Characteristics of Naive Bayes Classifier
- The Naive Bayes algorithm assumes that the predictors have independent and equal contributions in determining the output class.
- Naive Bayes model’s assumption that all predictors are independent of each other is not practical in real-world scenarios but still, this assumption gives a good result in most of the cases.
- Naive Bayes is commonly used for text classification where data dimensionality is often quite high.
Types of Naive Bayes Classifiers
There are 3 types of Naive Bayes Classifiers –
i) Gaussian Naive Bayes
This classifier is used when the values of predictors are continuous in nature and it is assumed that they follow Gaussian distribution.
ii) Bernoulli Naive Bayes
This classifier is used when the predictors are boolean in nature and it is assumed they follow Bernoulli distribution.
iii) Multinomial Naive Bayes
This classifier uses multinomial distribution and is mostly used for document or text classification problems
Gaussian Naive Bayes Classifier Example in Python Sklearn
In this section, we will take you through an end-to-end example of the Gaussian Naive Bayes classifier in Python Sklearn using a cancer dataset. We will be using the Gaussian Naive Bayes function of SKlearn i.e. GaussianNB() for our example.
1. Loading Initial Libraries
We will start by loading some initial libraries to load and visualize the dataset.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
2. Importing Dataset
Now we will upload the cancer detection dataset that we have obtained from Kaggle for performing our Naive Bayes classification.
dataset = pd.read_csv("datasets/cancer.csv")
3. Exploring Dataset
Let us take an initial look into the dataset with the help of head() function.
dataset.head()
| id | diagnosis | radius_mean | texture_mean | perimeter_mean | area_mean | smoothness_mean | compactness_mean | concavity_mean | concave points_mean | … | texture_worst | perimeter_worst | area_worst | smoothness_worst | compactness_worst | concavity_worst | concave points_worst | symmetry_worst | fractal_dimension_worst | Unnamed: 32 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 842302 | M | 17.99 | 10.38 | 122.80 | 1001.0 | 0.11840 | 0.27760 | 0.3001 | 0.14710 | … | 17.33 | 184.60 | 2019.0 | 0.1622 | 0.6656 | 0.7119 | 0.2654 | 0.4601 | 0.11890 | NaN |
| 1 | 842517 | M | 20.57 | 17.77 | 132.90 | 1326.0 | 0.08474 | 0.07864 | 0.0869 | 0.07017 | … | 23.41 | 158.80 | 1956.0 | 0.1238 | 0.1866 | 0.2416 | 0.1860 | 0.2750 | 0.08902 | NaN |
| 2 | 84300903 | M | 19.69 | 21.25 | 130.00 | 1203.0 | 0.10960 | 0.15990 | 0.1974 | 0.12790 | … | 25.53 | 152.50 | 1709.0 | 0.1444 | 0.4245 | 0.4504 | 0.2430 | 0.3613 | 0.08758 | NaN |
| 3 | 84348301 | M | 11.42 | 20.38 | 77.58 | 386.1 | 0.14250 | 0.28390 | 0.2414 | 0.10520 | … | 26.50 | 98.87 | 567.7 | 0.2098 | 0.8663 | 0.6869 | 0.2575 | 0.6638 | 0.17300 | NaN |
| 4 | 84358402 | M | 20.29 | 14.34 | 135.10 | 1297.0 | 0.10030 | 0.13280 | 0.1980 | 0.10430 | … | 16.67 | 152.20 | 1575.0 | 0.1374 | 0.2050 | 0.4000 | 0.1625 | 0.2364 | 0.07678 | NaN |
5 rows × 33 columns
Next, we will explore the columns present in the dataset through info() function.
dataset.info()
<class 'pandas.core.frame.DataFrame'> RangeIndex: 569 entries, 0 to 568 Data columns (total 33 columns): # Column Non-Null Count Dtype --- ------ -------------- ----- 0 id 569 non-null int64 1 diagnosis 569 non-null object 2 radius_mean 569 non-null float64 3 texture_mean 569 non-null float64 4 perimeter_mean 569 non-null float64 5 area_mean 569 non-null float64 6 smoothness_mean 569 non-null float64 7 compactness_mean 569 non-null float64 8 concavity_mean 569 non-null float64 9 concave points_mean 569 non-null float64 10 symmetry_mean 569 non-null float64 11 fractal_dimension_mean 569 non-null float64 12 radius_se 569 non-null float64 13 texture_se 569 non-null float64 14 perimeter_se 569 non-null float64 15 area_se 569 non-null float64 16 smoothness_se 569 non-null float64 17 compactness_se 569 non-null float64 18 concavity_se 569 non-null float64 19 concave points_se 569 non-null float64 20 symmetry_se 569 non-null float64 21 fractal_dimension_se 569 non-null float64 22 radius_worst 569 non-null float64 23 texture_worst 569 non-null float64 24 perimeter_worst 569 non-null float64 25 area_worst 569 non-null float64 26 smoothness_worst 569 non-null float64 27 compactness_worst 569 non-null float64 28 concavity_worst 569 non-null float64 29 concave points_worst 569 non-null float64 30 symmetry_worst 569 non-null float64 31 fractal_dimension_worst 569 non-null float64 32 Unnamed: 32 0 non-null float64 dtypes: float64(31), int64(1), object(1) memory usage: 146.8+ KB
From the above information, we can see that id and unnamed:32 columns are not useful, so we can remove them.
dataset = dataset.drop(["id"], axis = 1)
dataset = dataset.drop(["Unnamed: 32"], axis = 1)
3. Visualizing Dataset
contains information about patients who have a malignant (dangerous) tumor and the other half data for benign (safe) tumor.
Malignant Tumor Dataframe
M = dataset[dataset.diagnosis == "M"]
Benign Tumor Dataframe
B = dataset[dataset.diagnosis == "B"]
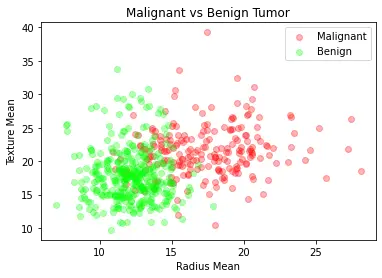
We will now visualize the malignant and benign tumors by looking at the mean radius and mean texture of these tumors.
plt.title("Malignant vs Benign Tumor")
plt.xlabel("Radius Mean")
plt.ylabel("Texture Mean")
plt.scatter(M.radius_mean, M.texture_mean, color = "red", label = "Malignant", alpha = 0.3)
plt.scatter(B.radius_mean, B.texture_mean, color = "lime", label = "Benign", alpha = 0.3)
plt.legend()
plt.show()
4. Preprocessing
Now we will be assigning a value of ‘1’ to malignant tumors and ‘0’ to benign tumors.
dataset.diagnosis = [1 if i == "M" else 0 for i in dataset.diagnosis]
We now break our dataframe into x and y. The x contains all the independent predictor variables and y contains the prediction of diagnosis.
x = dataset.drop(["diagnosis"], axis = 1)
y = dataset.diagnosis.values
5. Data Normalization
To improve the efficiency of the model it is always good to normalize data to bring them into a common scale.
# Normalization:
x = (x - np.min(x)) / (np.max(x) - np.min(x))
6. Test Train Split
Now with the help of sklearn library train_test_split module, we will split the dataset into training and testing parts.
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size = 0.3, random_state = 42)
7. Sklearn Gaussian Naive Bayes Model
Now we will import the Gaussian Naive Bayes module of SKlearn GaussianNB and create an instance of it. We can pass x_train and y_train to fit the model.
from sklearn.naive_bayes import GaussianNB
nb = GaussianNB()
nb.fit(x_train, y_train)
GaussianNB()
8. Accuracy
The following accuracy score on the test data shows how well our model Sklearn Gaussian Naive Bayes model has performed for predicting cancer.
print("Naive Bayes score: ",nb.score(x_test, y_test))
Naive Bayes score: 0.935672514619883
- Also Read – Linear Regression in Python Sklearn with Example
- Also Read – Python Sklearn Logistic Regression Tutorial with Example
Conclusion
Coming to the end of the article, where we explain Naive Bayes and showed you an end-to-end example of implementing Gaussian Naive Bayes in Sklearn with Cancer dataset. Hope this small introductory tutorial was quite helpful for you.
Reference – https://www.kdnuggets.com/2020/06/naive-bayes-algorithm-everything.html
-
I am Palash Sharma, an undergraduate student who loves to explore and garner in-depth knowledge in the fields like Artificial Intelligence and Machine Learning. I am captivated by the wonders these fields have produced with their novel implementations. With this, I have a desire to share my knowledge with others in all my capacity.
View all posts


